Sample Requirements, Submission, & Data
Library Design Requirements

If you are new to Deep Sequencing, see the Resources section for basic information and helpful links.
The diagram below shows the basic structure of an Illumina library and the terms used on our submission ticket to describe its parts.
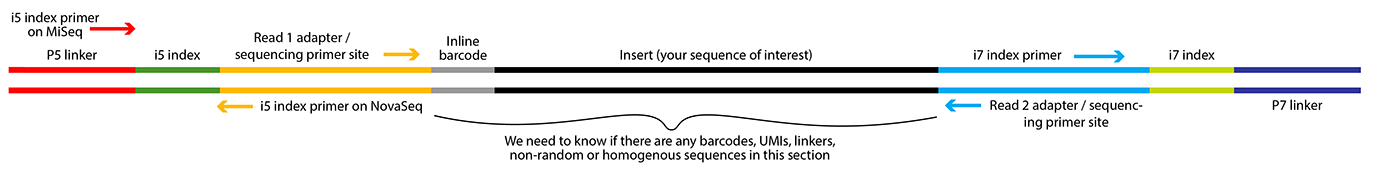
Please fill out the service ticket(s) as completely and accurately as you can. It is important that we receive the correct information about the primers and adapters used to build your library. Not all library types will run on all instruments or can be read in both directions.
- For premade/completed libraries:
Submit a Sequencing Ticket AND a Sample Information Sheet (paper + electronic). - For library prep from DNA, cDNA, RNA, or cells:
Submit a Library Prep Ticket, a Sequencing Ticket AND a Sample Information Sheet (paper + electronic) with as much information filled out as possible. Include the number of cells submitted for each sample if applicable. - For library prep from a NanoString collection:
Submit a Library Prep Ticket, the _SeqCodeIndices.csv file from the NanoString instrument (paper + electronic), AND a Sequencing Ticket. - If you are an internal UMass customer, the Core will accept an email from your PI authorizing charges in lieu of a signature.
- For external users: please submit an External (non-UMass) customer mandatory information form in addition to the above tickets.
Indexing Considerations:
- Single and Dual Multiplexing are distinct from Single and Paired-End reads. Single multiplexed libraries have one index (i7) at the P7 end, while dual multiplexed libraries have indexes at BOTH the P7 and P5 ends (i7 and i5). Each index that is placed in an adapter requires a separate read that is distinct from the insert reads (see diagram above).
- Dual Indexing or Unique Dual Indexing with AT LEAST 8-base indices is required for very high throughput sequencing, such as with the NovaSeq. The Core strongly recommends using Unique Dual indices for better base balance.
- For more details on multiplexing, please see the Indexing section on our Resources page.
Custom designs and controls:
- A good balance of the four bases is important for accurate base-calling. If you have used non-random sequence such as inline barcodes, linkers, spike-in controls, etc., please inform us on the relevant section of the ticket. These additions to the insert often require a sequencing control to improve base-balance.
- If you have added a spike-in control to your sequencing library, please be sure to specify that information on the ticket. Controls for normalization ("Active Motif", other antibody spike-ins) often have poor base-balance/repetitive motifs, and can badly overseed on a flowcell.
- If the kit or method you used for sample preparation does not work with the standard Illumina primer mix, please check off 'custom adapters' on the ticket and make arrangements for the Core to be provided with the custom sequencing primer. This must be pre-approved if you intend to run on the NovaSeq.
- If you are doing a custom design to include adapters, your own index scheme, or custom sequencing primers, these sequences must span the P5 and P7 attachment sequences on either end of the library construct and be compatible with the instrument you intend to run on. If you do not agree to submit the sequence for verification, then you will be run with only the primers you request and at your own risk.
For those preparing their own libraries, we reserve the choice to hold back a sample which fails QC and contact you to discuss options. The use of a carrier agent during the final stages of workup is discouraged. These can interfere with cluster formation and reduce the number of reads generated during sequencing.
Downloadable Forms and Documents

NovaSeq/MiSeq Analysis Ticket 2025 and Sample Information Sheet 2025 (You must complete BOTH for each GROUP of samples that will run together! You do not need to fill out a ticket for each sample in a mix.)
Online NovaSeq/MiSeq Analysis Ticket. Please first download a blank Sample Information Sheet above and fill in the information requested to have it ready for upload. Your PI will receive an automated email and must respond to confirm in lieu of a signature.
External (non-UMass) customer mandatory information form ( Intake Form )
Library Construction Order Form 2025 (Please fill out one for each GROUP of samples that will be mixed together. Add the Sequencing Analysis Ticket and Sample Information Sheet above for sequencing requests.)
Sample Submission Instructions ( click here for submission instructions )
DSCL workflow diagram ( Workflow pdf )
Sample Submission Process

Please make sure sample tickets (paper or electronic) are submitted at the same time as or prior to your sample submission. We need to know the correct workflow for your samples before they can be processed.
Libraries should be submitted at 10nM concentration in 25μl of ultrapure water or EB. If your library doesn't have that high a concentration, don't panic! The Core will accept samples with concentrations as low as 2nM, depending on library quality at QC. Libraries below 2nM in concentration will only be run "at-your-own-risk", and outcome will not be guaranteed by the Core.
Please use 1.5mL tubes, 0.5-0.65mL tubes, or PCR strips that are labeled on the top of the tube. Do not submit single PCR tubes, and avoid long labeling strips that would prevent the tube from fitting in a centrifuge. In the Illumina pipeline, sample names are limited to 20 characters, which can include alphanumerics, "-", or "_".
Please note that a sample submitted for sequencing is entered in the workflow and queue, which engages time and services. If you want the QC analysis only for your own information and are not certain you wish to move forward, please use the MBCL Fragment Analyzer or other QC methods (e.g. Topo Cloning, Gel Purification).
Typically we suggest that you send indexed libraries separately and let us do the mixing by molarity based on our internal QC. This helps with any potential troubleshooting if some samples in the mix underperform. However, if you prefer to premix your samples or choose a different mixing method (such as by volume), you are welcome to.
Drop-off locations:
Samples can be left in the labeled fridges at these locations
for 11:00 AM pickup each business day:
AS8-2016
N2 freezer bay near the main lobby elevators
LRB 6th floor mailroom
We are also available by appointment if you need to visit with us.
Our shipping address for off-campus customers is
Rose-Gordon Bldg. Rm. 141, 222 Maple Ave, Shrewsbury, MA, 01545.
If you are shipping your sample(s), please do not ship for a Friday delivery! Carriers may not deliver them in time to be properly stored before the weekend.
Sequencing Process

The Core Lab staff will QC your submitted library and enter it in the sequencing queue. The Core will email you a copy of the QC results upon request. If you then choose not to move forward with sequencing, the Core will bill for the QC and processing to recoup costs.
Samples will be processed on a first-come, first-served basis. For runs involving multi-lane flowcells, this may include wait time until the run type is full. See our FAQ section for frequently asked questions about queue times.
You will receive an email notification when your data is ready. Instructions on how to retrieve your data will be included in the email.
Data Transfer

The amount of data generated from a high-throughput run can be extremely large. We recommend users get a Globus account. You do not need a full Globus subscription to receive data; however you do need a registered email account. Information about Globus subscription and personal use is available at https://app.globus.org.
If you need an alternative data delivery method, please contact us to discuss it prior to submitting samples.
- Please note:
- You will get an email notification when your data files are ready.
- If you require anything besides the fastq files, you must let us know at the time you submit your sample.
- Data files will be deleted 5 business days after notification; please backup/copy/archive your data in your own workspace.
- The DSCL does not have the resources to archive or recover data. Please ensure that you safely back your data up!
Data Analysis

Please see the DeepSeq Data section for information about data analysis. If you would like to consult with an applications specialist, please email us to get on the schedule.
IF THERE IS A PROBLEM WITH YOUR RUN OR QUESTIONS ABOUT YOUR DATA, please let us know as soon as possible. The run metrics and machine files cannot be held for long because they are extremely large. The sooner we know there is a problem, the more likely we'll be able to help easily. In order to help sort out a problem we may ask a lot of questions and details about your sample and your analysis methods. This information is required in order for us to engage the tech support systems provided by our vendors.
Please note that if the control lane on the run failed in any way, the entire flow cell is rerun. If your data is delivered to you, then the controls all passed spec., which means that the instrument, reagents, and chemistry functioned properly. The Core will then begin investigating other issues. These may include sample design, base balance, sizing, indexing, or degraded DNA. They may also include computational and analysis issues with the pipeline.
We'll do whatever we can to get things working, but the more information we have, the better. You can find a troubleshooting guide for libraries here which is updated as new information is gathered.
Pricing and Payment Policy

Please email DeepSequencingCoreLabs@umassmed.edu to get a quote for your desired services.
Processing and analyzing a sample requires time and reagents. Payment for services is the responsibility of the user submitting the sample and should be rendered in a timely fashion. In the event of reagent or equipment failure, samples will be rerun at the next possible opportunity with no additional charge.
Clients withdrawing samples prior to the analysis run will be charged a fee to recover QC assay costs. For the return of archived post-analysis samples, clients will be charged a delivery fee per sample.
A signed ticket or email authorization indicates that the holder of the listed account consents to payment in accordance with this policy.



